AP Biology 6.2 - Replication
In this section, we will focus on how DNA is replicated – an essential process for life on Earth. We’ll start by looking at the various theories of DNA replication that were originally proposed after the structure of DNA was understood: conservative replication, semiconservative replication, and dispersive replication. The Meselson-Stahl experiment famously determined that semiconservative replication was the most accurate theory. After this, we’ll dive into the process of DNA replication. We’ll start with the helicase and topoisomerase enzymes that unwind the double-stranded DNA molecule. After the first quiz, we’ll see how DNA polymerase actually functions to build a new DNA strand from the template strand. Lastly, we’ll see the final steps in DNA replication that finish off the process and how DNA replication is very similar to RNA transcription.
Video Tutorial
The following video summarizes the most important aspects of this topic!
To watch more tutorial videos like this, please click here to see our full Youtube Channel!
Resources for this Standard
For Students & Teachers
- Overview & Video Tutorial (This article)
- Quick Test Prep
- Crossword Puzzle
For Teachers Only
ENDURING UNDERSTANDING
IST-1
Heritable information provides for continuity of life.
LEARNING OBJECTIVE
IST-1.M
Describe the mechanisms by which genetic information is copied for transmission between generations.
ESSENTIAL KNOWLEDGE
IST-1.M.1
DNA replication ensures continuity of hereditary information–
- DNA is synthesized in the 5’ to 3’ direction.
- Replication is a semiconservative process — that is, one strand of DNA serves as the template for a new strand of complementary DNA.
- Helicase unwinds the DNA strands.
- Topoisomerase relaxes supercoiling in front of the replication fork.
- DNA polymerase requires RNA primers to initiate DNA synthesis.
- DNA polymerase synthesizes new strands of DNA continuously on the leading strand and discontinuously on the lagging strand.
- Ligase joins the fragments on the lagging strand.
EXCLUSION STATEMENT
The names of the steps and particular enzymes involved — beyond DNA polymerase, ligase, RNA polymerase, helicase, and topoisomerase — are beyond the scope of the course and the AP Exam.
6.2 Replication Overview
The process of DNA replication is a lot like playing with Lego blocks. If you are trying to build a specific structure, the pieces have to be carefully added in just the right order. In this metaphor, your hands are like enzymes – carefully arranging and joining the Lego blocks (representing nucleotides) in a specific order. This is the only way you can start with a pile of blocks and end with a functioning toy.
DNA replication is a convoluted process, with many moving parts and pieces that need to be combined in a specific order. Yet, somehow this process happens in your body every day – with almost no errors. Several proteins, RNA primers, and other molecules work together to unwind a DNA molecule and replicate it exactly in a semiconservative way. This will definitely be on the AP test. So, stick with us as we cover everything you need to know about DNA replication!
Once the structure of DNA was fully understood, the process of DNA replication still had to be determined. Several theories were presented as to how DNA was replicated and passed into daughter cells after mitosis or binary fission.
The first theory, conservative replication, would happen if both strands of the template DNA molecule stayed bonded and intact while serving as a template for a brand new and identical double-stranded DNA molecule. The second theory, semiconservative replication, postulates that the two original strands of DNA separate, each serving as a template for a new strand – creating DNA molecules that have one original strand and one new strand. The third theory, dispersive replication, considered that in order for strands not to become twisted during replication, the original strand would need to be cut every 10 nucleotides or so. This would create replicated molecules with segments of new double-stranded DNA and segments of original double-stranded DNA.
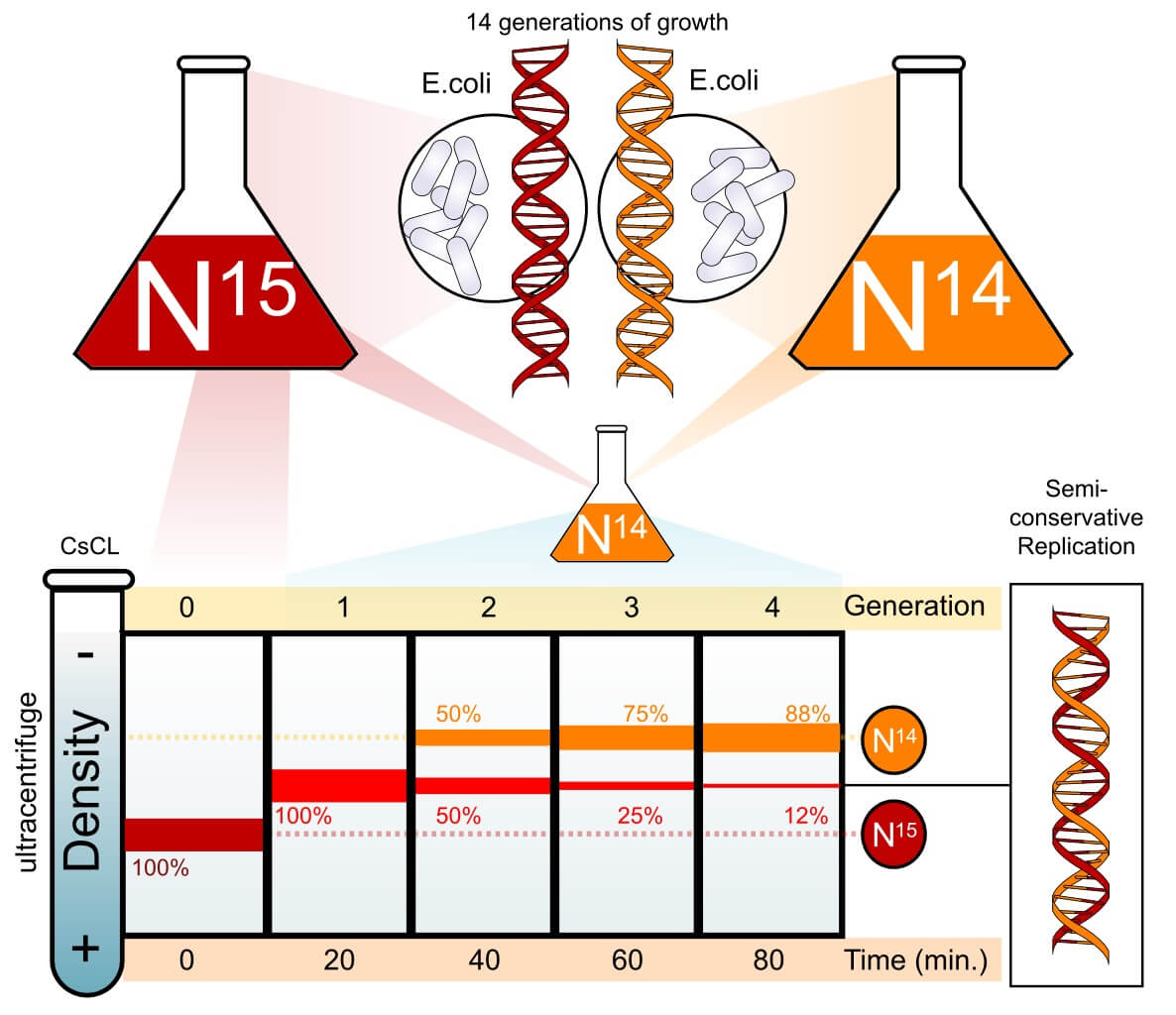
In order to determine which of these theories were correct, a fantastic experiment was devised. The Meselson-Stahl experiment started with the observation that E. coli bacteria use the elements in their environment to construct new nucleotides and build new DNA molecules. Thus, if you place these bacteria in a solution with normal nitrogen-14, the DNA will be slightly less dense than if you place the bacteria in a solution with the isotope nitrogen-15. You can literally measure the difference in their densities by centrifuging the two types of DNA at high speeds. The DNA with N14 will appear higher in the centrifuged test tube than the DNA with N15.
So, the researchers started by growing E. coli in a solution with heavy nitrogen until all the DNA in this colony was made from nitrogen-15. Then, this colony of bacteria was transferred to a growing medium that contained only nitrogen-14. This is what they found as they centrifuged and measured the DNA collected from every generation of bacteria. The original generation showed a single band at the nitrogen-15 level. However, the first generation of bacteria showed a single band at an intermediate density – between nitrogen-14 and nitrogen-15 levels.
Since conservative replication would have produced two distinct strands – half of the DNA would have been heavy, while and the new half would have been entirely light – this first generation eliminated the conservative hypothesis. But, to determine which of the other two hypotheses were correct, researchers had to continue the experiment. When researchers allowed the bacteria to reproduce a second time, they found two distinct bands – one entirely light and the other intermediate. Continuing for several more generations, the percentage of intermediate DNA slowly decreased until only nitrogen-14 DNA was left.
Since the second generation had an entirely lightweight band of DNA, dispersive replication was also ruled out. Dispersive replication would have only ever produced one band that slowly became closer and closer to lightweight over time. Therefore, Meselson and Stahl were able to show conclusively that DNA replication is a semiconservative process.
Thanks to Meselson and Stahl, we know that a semiconservative process takes place that replicates the DNA. But, in order for this process to take place, the DNA strands must first unwind. It took researchers several more decades to figure out how this process worked since microscopes only go so far and there were a few obvious hurdles to overcome in semiconservative replication.
First, the two strands of DNA must be separated. This is the first problem. But, if you simply grab the two strands and pull them apart, the twists that are naturally present in the two strands will be pushed into the remaining double-stranded section, creating supercoiled DNA that knots-up on itself. Supercoiling is similar to what happens when you are trying to get the knots out of your headphones. If you simply grab the two headphones and pull on them, you end up creating tight knots that are hard to untangle.
To overcome these two hurdles, the process of DNA replication uses two very important enzymes: helicase and topoisomerase. Each of these enzymes is absolutely necessary for the processes of DNA replication and RNA transcription.
Helicase is an ATP-powered enzyme that pushes apart the two strands of DNA. As helicase encounters a DNA molecule, regions of the helicase enzyme interact with individual nucleotide bases, destabilizing the hydrogen bonds present. As ATP is supplied to the helicase molecule, it pulls on each of the separated strands, separating the base pairs as it travels along the DNA molecule. This allows DNA polymerase to access each strand and start the replication process. Keep in mind that there are many different types of helicase enzymes that act slightly differently in different organisms and situations.
However, the cell still needs to deal with the problem of supercoiling. For this, the enzyme topoisomerase is used. You can remember this because “topo” is in reference to DNA topology, which is the tertiary, 3D shape of the DNA – like a topographic map. Topoisomerase has this name because its job is to change the 3D shape of DNA. Specifically, topoisomerase will relieve the torsion that has been created as helicase separates the strands of DNA. To do this, topoisomerase will cut the sugar-phosphate backbone. The torsion will unwind from the DNA strand, giving it back its normal twist and preventing any supercoiling or knotting that would otherwise occur. Then, topoisomerase reconnects both strands in their original position.
Topoisomerase will continue moving ahead of the helicase enzyme as the DNA is replicated to continually relieve the torsion as it builds.
After the strands of DNA have been opened up, DNA polymerase can come in and get to work. But, DNA polymerase cannot just latch on anywhere and start the process of replication. It needs a little help to get started.
First, a protein called primase comes in and adds an RNA primer to the exposed template strand. This gives DNA polymerase an exposed 3’ hydroxyl group that it can begin adding new nucleotides to. Using free nucleotides from the cytosol, DNA polymerase ensures base pairing occurs between the free nucleotides and the existing template. Then, it catalyzes the formation of the phosphodiester bond between these nucleotides.
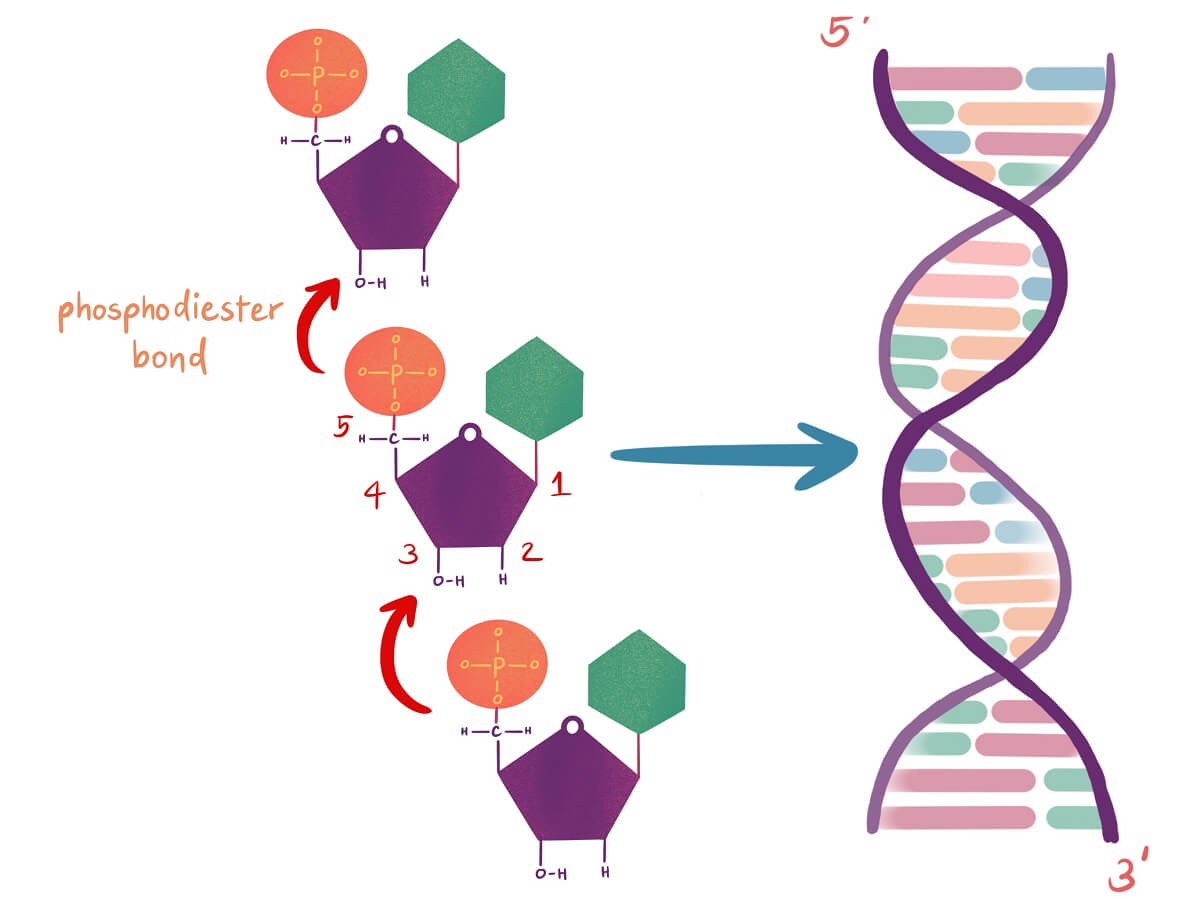
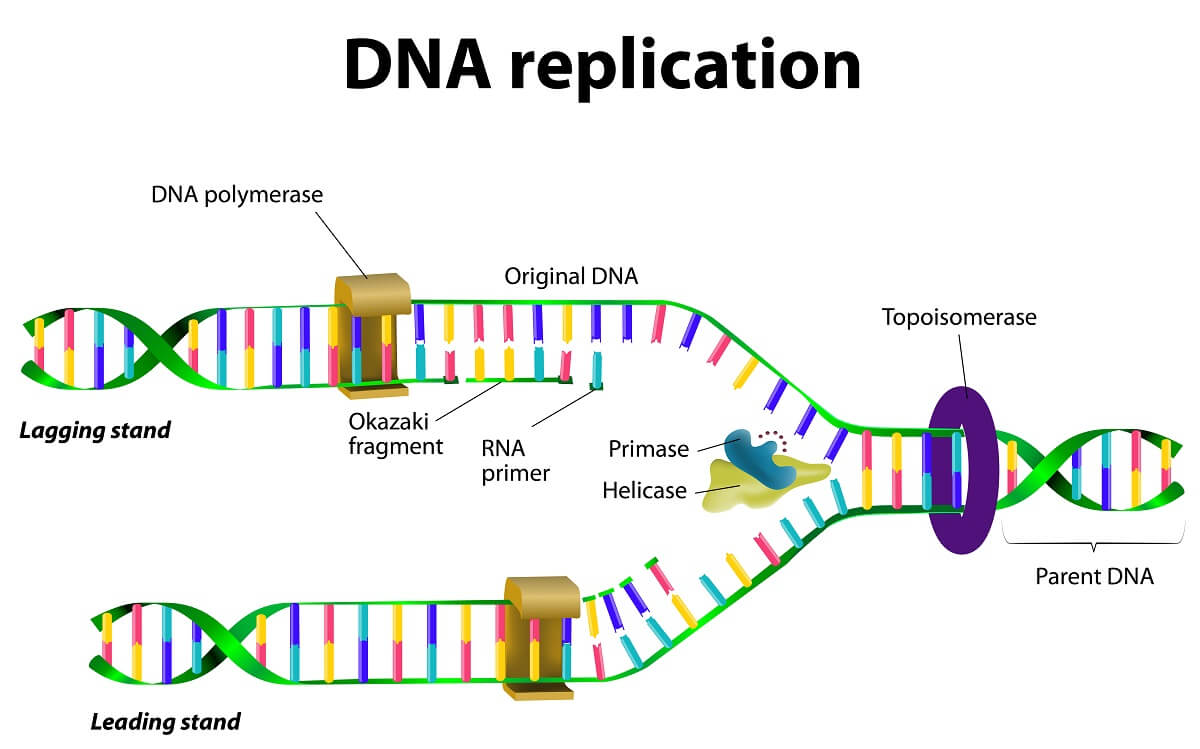
The DNA polymerase can only add nucleotides in the 5’ to 3’ direction. In other words, the DNA polymerase can only catalyze the bond between a 3’ OH group and the phosphate group attached to the 5’ carbon of the new nucleotide. This creates a difference between the two different strands of DNA being replicated that is important to understand.
Since the two strands of DNA run antiparallel, the two DNA polymerase molecules operate in different directions. The leading strand is created in the same direction that helicase is opening up the DNA, while the lagging strand is created in the opposite direction. While the DNA polymerase on the leading strand can add nucleotides continuously as helicase opens up new portions of the strand, the DNA polymerase on the lagging strand must continuously reattach to the newly opened template strand. This creates a large number of Okazaki fragments instead of a single new strand. In the next slide, we’ll see how these fragments are connected.
You may have noticed that for DNA polymerase to attach to the DNA template strand, it needs an RNA primer to latch onto. These must be removed and filled in with DNA nucleotides of the process of DNA replication to be complete.
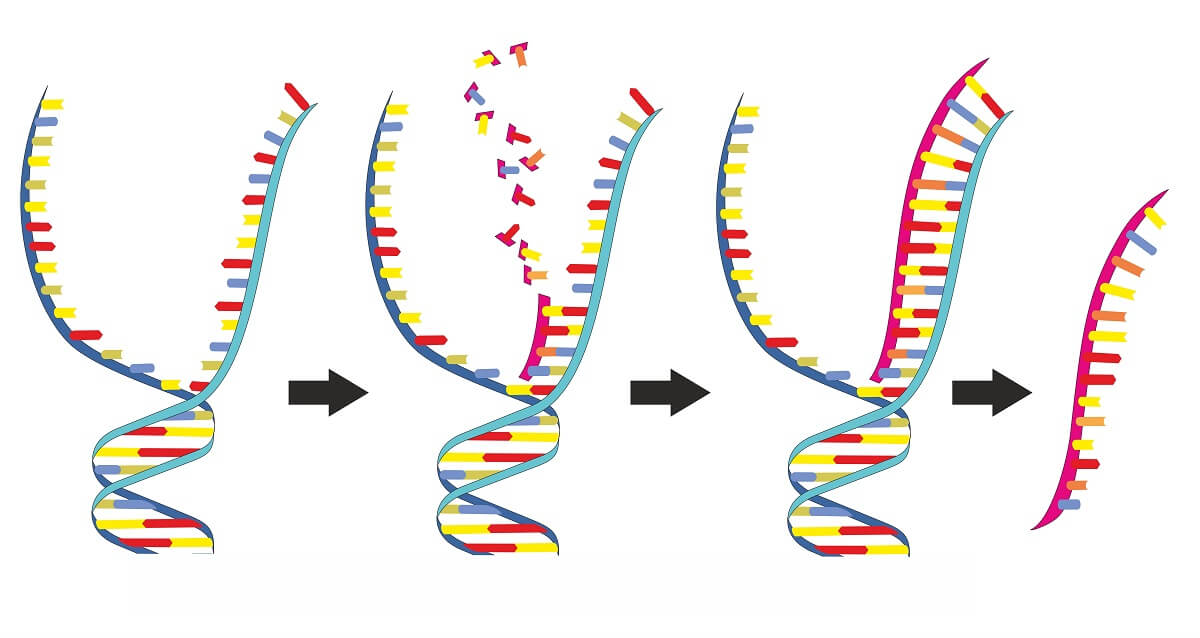
To do this, another enzyme (called RNase) is needed to first break down the RNA primers. This leaves a small gap in the DNA strand that must be filled in. The final enzyme that carries out the process of filling in all the gaps where RNA primers sat and fully connecting the newly synthesized DNA strand is ligase.
Free-floating nucleotides in the cytosol fill in the gap by base-pairing with the nucleotides on the template strand. Ligase works by using ATP to create the phosphodiester bonds to fully complete the sugar-phosphate backbone. This leaves the new DNA molecule fully intact and completed!
Keep in mind that the lagging strand will require the services of ligase much more than the leading strand – since each Okazaki fragment begins with an RNA primer. Each of these primers must be replaced, filled, and the backbone fully ligated by ligase. Also keep in mind that DNA ligase enzymes work in other situations as well, such as when scientists add foreign DNA to a plasmid to create recombinant DNA and fully bond the new pieces in place.
You should also remember that many of the same enzymes discussed in DNA replication are used in RNA transcription.
